Hi Open3D community,
I am quite new here, I’d like someone to show me around based on my code 
I am trying to reconstruct a surface from (few) experimental data. I’d like to have as much flexibility/tunability as possible.
My code is something like this:
import open3d as o3d
sys.path.append('..')
output_path=(r"C:\Users\Giammarco\Desktop\PYTHON_graphs\OUTPUTS\\")
poisson_mesh=[]
densities=[]
pcd = o3d.geometry.PointCloud()
pcd.normals = o3d.utility.Vector3dVector(np.zeros((1, 3))) # invalidate existing normals
#load the point cloud
point_cloud=np.array([x,y,z]).T
cloud = PyntCloud.from_instance("open3d", pcd)
pcd.points = o3d.utility.Vector3dVector(point_cloud)
#resise the scale of the sample
vox_grid = o3d.geometry.VoxelGrid.create_from_point_cloud(pcd, 1.)
#presetn in all approaches of plc
kdtree = cloud.add_structure("kdtree")
testc = cloud.get_neighbors(k=5)
distances = pcd.compute_nearest_neighbor_distance()
avg_dist = np.mean(distances)
#compute the normals
pcd.estimate_normals(); #mandatory
#orient the normals
#Number of nearest neighbours: 5 is the minimum to have a closed surface with scale >= 2
pcd.orient_normals_consistent_tangent_plane(7)
#Poisson algorithm
poisson_mesh, densities = o3d.geometry.TriangleMesh.create_from_point_cloud_poisson(pcd, depth=9, width=0, scale=3.5, linear_fit=False)
bbox = pcd.get_axis_aligned_bounding_box()
p_mesh_crop = poisson_mesh.crop(bbox)
# cleaning
# p_mesh_crop =poisson_mesh.simplify_quadric_decimation(6000)
# p_mesh_crop.remove_unreferenced_vertices
# p_mesh_crop.remove_degenerate_triangles()
# p_mesh_crop.remove_duplicated_triangles()
# p_mesh_crop.remove_duplicated_vertices()
# p_mesh_crop.remove_non_manifold_edges()
#designing the surface colour
#densities are the real density of features
densities = np.asarray(densities)
density_colors = plt.get_cmap('viridis')((dgo - dgo.min()) / (dgo.max() - dgo.min()))
density_colors = density_colors[:, :3]
#works for the plotting in o3d
poisson_mesh.vertex_colors = o3d.utility.Vector3dVector(density_colors)
o3d.io.write_triangle_mesh(output_path+"bpa_mesh.ply", dec_mesh);
o3d.io.write_triangle_mesh(output_path+"p_mesh_c.ply", poisson_mesh);
# o3d.io.write_triangle_mesh(output_path+"p_mesh_c.ply", p_mesh_crop);
# my_lods = lod_mesh_export(p_mesh_crop, [100000,50000,10000,1000,100], ".ply", output_path)
my_lods = lod_mesh_export(poisson_mesh, [100000,50000,10000,1000,100], ".ply", output_path)
# o3d.visualization.draw_geometries([pcd, p_mesh_crop], mesh_show_back_face=True)
# o3d.visualization.draw_geometries([pcd, poisson_mesh],mesh_show_back_face=True)
# o3d.visualization.draw_geometries([pcd, poisson_mesh[100000]],point_show_normal=True)
# tri_mesh_pois.show()#designing the surface colour
#densities are the real density of features
densities = np.asarray(densities)
density_colors = plt.get_cmap('viridis')((dgo - dgo.min()) / (dgo.max() - dgo.min()))
density_colors = density_colors[:, :3]
#works for the plotting in o3d
poisson_mesh.vertex_colors = o3d.utility.Vector3dVector(density_colors)
o3d.io.write_triangle_mesh(output_path+"p_mesh_c.ply", poisson_mesh);
my_lods = lod_mesh_export(poisson_mesh, [100000,50000,10000,1000,100], ".ply", output_path)
#SHORTCUTS from keyboard: n = show normals, q = quit, w = mesh
o3d.visualization.draw_geometries([pcd, poisson_mesh],mesh_show_back_face=True)
output:
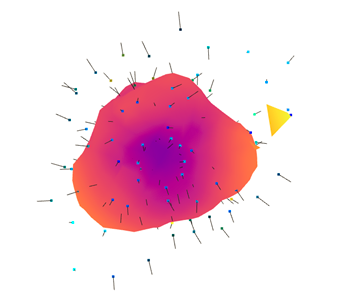
-
I am concern about the
create_from_point_cloud_poissonfit model option: is there a way to tune its parameters more than justdepthandsize? Is there an iterative process that I should set up for better conversion (e.g. threshold)? As you can see the distance btw the calculated surface and the experimental point is quite big. -
Is the estimation of normals properly set up? In the second output, some directions are still quite random.
I tried this syntax too:pcd.estimate_normals(search_param=o3d.geometry.KDTreeSearchParamHybrid(radius=0.05,max_nn=20));, but It doesn’t converge to a closed surface, just a plane (I am limited to one picute per post).
Please, give me feedbacks on my code and suggestions on how to improve it.
Thank you for your support!
